SEVA: Building the Backbone of European Synthetic Biology
Synthetic biology is rapidly gaining traction for its contributions to health, climate and industrial biotechnology. This progress has relied on the development of open-access tools that make research reproducible and transferable. One of Europe’s most important contributions to synthetic biology infrastructure is the Standard European Vector Architecture (SEVA) web-based resource and plasmid vector repository.
We spoke with Esteban Martínez, SEVA co-founder and researcher at the National Centre for Biotechnology (CNB-CSIC) in Madrid, about how SEVA started, how it has since evolved, and where it is heading in the age of AI and automation.
From a lab problem to a community resource
Prior to the conception of SEVA, Martínez and colleagues in Victor de Lorenzo’s lab encountered practical challenges while working with the environmental bacterium Pseudomonas putida, a non-model organism. Although this bacterium showed strong biotechnological potential, the available genetic tools were limited.
Martínez explains that the lab “didn’t have the proper tools to be able to do all the things that we wanted to do”. Plasmids designed for use in non-model organisms were scarce, and adapting existing systems was laborious. Rather than continually adapting incompatible systems, the team began developing a modular collection that allows easy exchange of genetic elements, facilitating the creation of plasmids with desired properties and novel functionalities.
Initially, this collection was simply meant to serve their own lab, but when it was first shared publicly in 2013 external researchers were eager to contribute. The database expanded rapidly, leading the team to repeatedly adapt the plasmid nomenclature to account for the growing number of biological parts included. An update on the most recent iteration, SEVA 4.0, was published in 2023.
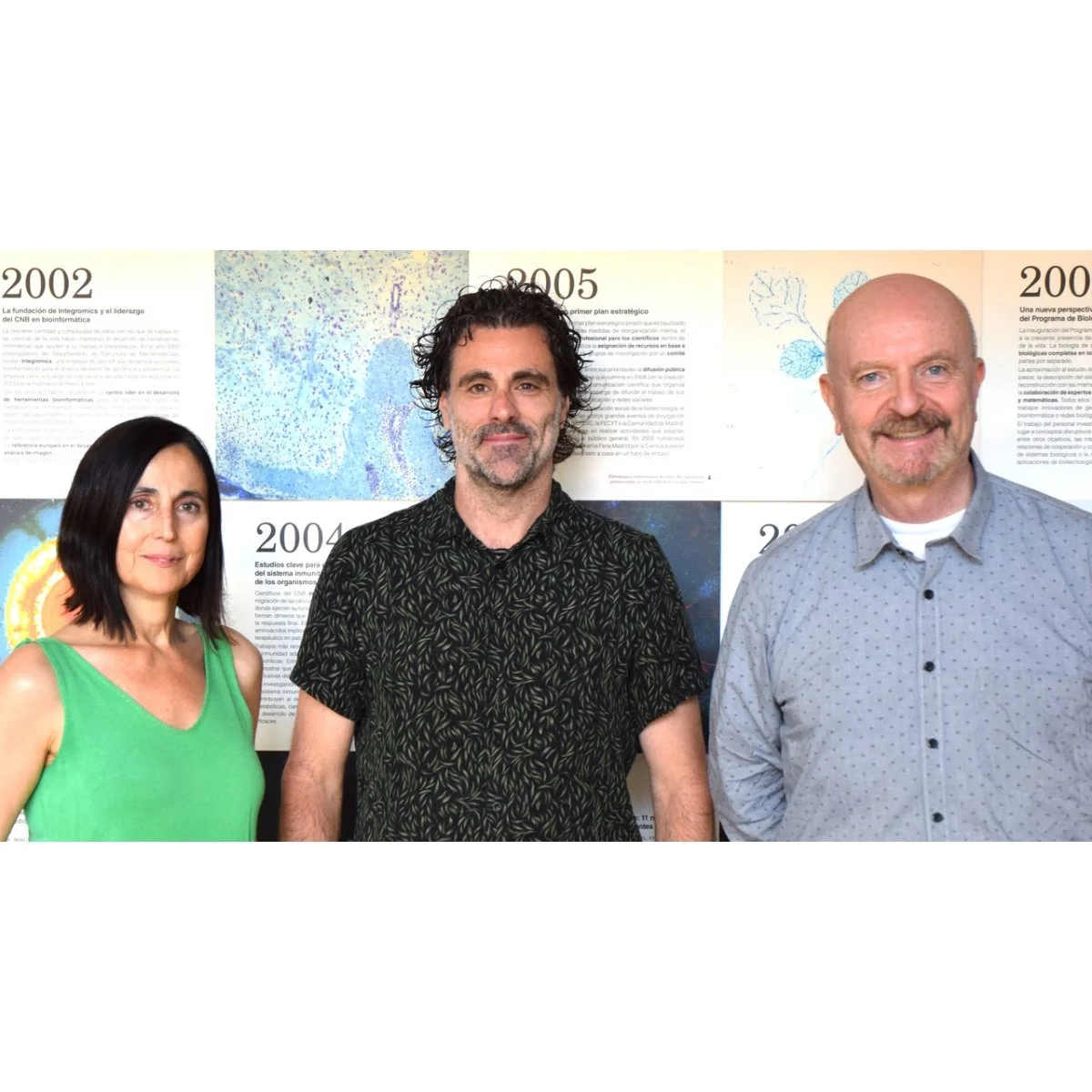
Figure 1. The SEVA Team. Left to right: Sofía Fraile, Esteban Martínez, Víctor de Lorenzo. Image Credit: CNB-CSIC.
Today, SEVA has distributed more than 5,000 plasmids to researchers in 48 countries. In 2025 alone, over 500 canonical SEVA plasmids were shipped worldwide. Despite the scale of this operation, the core team that maintains the collection consists of just three people. Martínez, Víctor de Lorenzo, and Sofía Fraile manage SEVA alongside their full-time research and academic posts.
The collection has grown not only through internal efforts but also through contributions from collaborators across Europe, including members of the “SEVA Commonwealth”. Labs that develop standardised modules used in SEVA plasmids are encouraged to send them to the collection so others can benefit.
SEVA operates as a fully open-access resource. Plasmids are provided free of charge to researchers worldwide. The sequences are openly available, and recipients are not restricted in their use.
The four rules of SEVA
SEVA’s strength and utility lies in its strict standardisation framework. Plasmids in the canonical SEVA collection follow four key rules:
Use of the minimal functional DNA sequence for genetic modules
Removal of common restriction sites from each module to facilitate easy module exchange
Flanking of origin of replication, antibiotic markers, and functional (cargo) modules with defined restriction sites
Use of a structured nomenclature system where each canonical plasmid carries the suffix “pSEVA” followed by four numbers encoding the antibiotic resistance marker, origin of replication, cargo, and an optional additional “gadget” module (modules that confer new utility to the basic SEVA backbone).
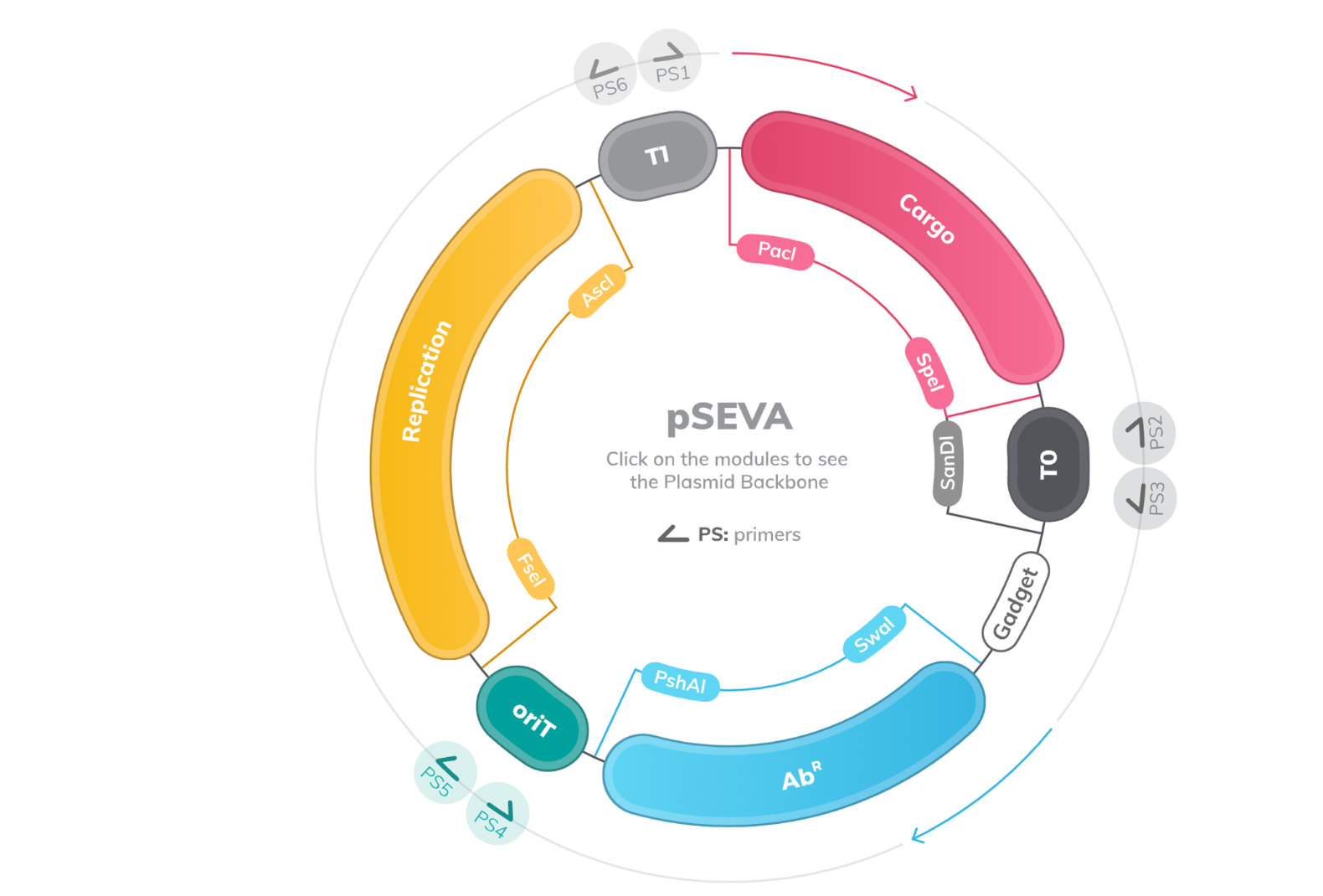
Figure 2. The canonical SEVA plasmid backbone. An interactive view for exploring the module features is available on the SEVA website (image used with permission from seva-plasmids.com).
These rules streamline module exchange and reduce unnecessary sequence complexity. By eliminating common restriction sites and standardising module boundaries, researchers can easily swap components without implementing new designs.
Plasmid standardisation requires effort and resources. Martínez acknowledges that “not all laboratories are able to invest the time and money to create a standard plasmid. They just want the plasmid to work.” To address this, the database includes “SEVA siblings”: plasmids that use the SEVA backbone but do not comply with all four rules. pSEVA-sibs are available to browse through the SEVA website and may later be standardised by collaborators or volunteers.
Expanding biotechnology beyond model organisms
A major motivation behind SEVA was to expand genetic tools beyond E. coli. While E. coli remains the dominant laboratory workhorse, many applications require alternative hosts.
SEVA enables researchers working with environmental or non-model Gram-negative bacteria to systematically test different origins of replication, antibiotic resistance markers, cargoes and gadgets of interest. Further, SEVA plasmids include an origin of tranfer (oriT) sequence, allowing transfer by conjugation, thus increasing their versatility across species.
Collaborations have enabled the expansion of the SEVA collection beyond Pseudomonas. For example, Felipe Lombó’s lab also faced problems resulting from limited tool availability for biotechnological research using Streptomyces species. Collaborating enabled the design of canonical SEVA plasmids for Streptomyces species, yielding 23 novel shuttle vectors and a novel selection marker. Expansion of the SEVA database relies on partnerships, with plasmid validation in new organisms driven by labs with relevant expertise.
The importance of standardisation for AI and automation
The modular design of SEVA plasmids creates large combinatorial potential. Martínez notes that “in the SEVA collection, we have a lot of different modules. If you combine all of them, it’s like a combinatorial explosion” yielding tens of thousands of possible plasmids. The SEVA framework allows labs to use the SEVA backbone to generate a plasmid with a combination of modules particularly suited to their application.
One of SEVA’s immediate goals is to automate plasmid storage and shipment, intending to integrate the collection with an upcoming biofoundry at CNB-CSIC. Given the growing demand and small team size, automation could significantly improve sustainability and allow further growth of the collection.
Beyond plasmid architecture, Martínez highlights a broader need in synthetic biology: the standardisation of measurement and data. He emphasises the importance of metrology and the adoption of consistent methods for characterising gene expression, fluorescence and other biological outputs.
Without comparable measurement standards, data generated in different laboratories remain difficult to integrate. This limits reproducibility and hampers efforts to build predictive computational models.
Martínez argues that wider adoption of standards will strengthen synthetic biology. Increased use of Synthetic Biology Open Language (SBOL) for standardised biological part description, Synthetic Biology Markup Language (SBML) for software and modelling formats, and biobanking practices would improve consistency across the field. Martínez believes “once we start properly characterizing biological parts, we could use those data to train algorithms to feed artificial intelligence [..] to predict how to assemble different genetic constructs and predict if it's going to work or not”.
He notes that generating high-quality characterisation data requires time and sustained effort. While foundational tool-building may not always be viewed as “exciting,” it lays the groundwork for developing ambitious synthetic biology projects.
Building a European infrastructure for synthetic biology
SEVA illustrates how long-term investment in standardised tools can help shape an ecosystem. By focusing on modularity, reproducibility and open access, the initiative has helped expand synthetic biology beyond traditional model organisms while strengthening collaboration within Europe and the wider global community.
As automation, biofoundries and computational design become increasingly central to synthetic biology, infrastructures like SEVA provide the standardised tools necessary for scalable innovation in synthetic biology, biotechnology & beyond.